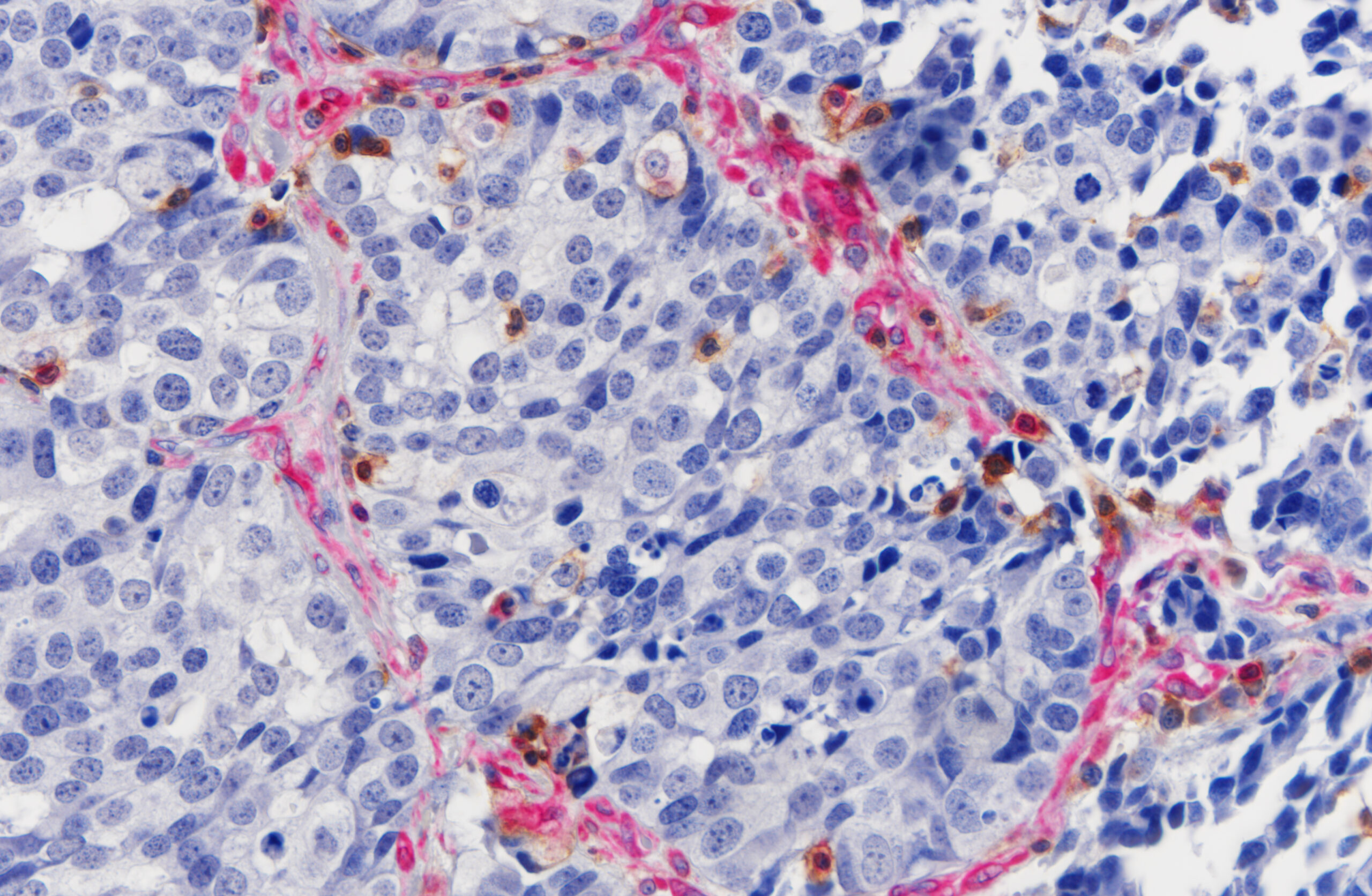
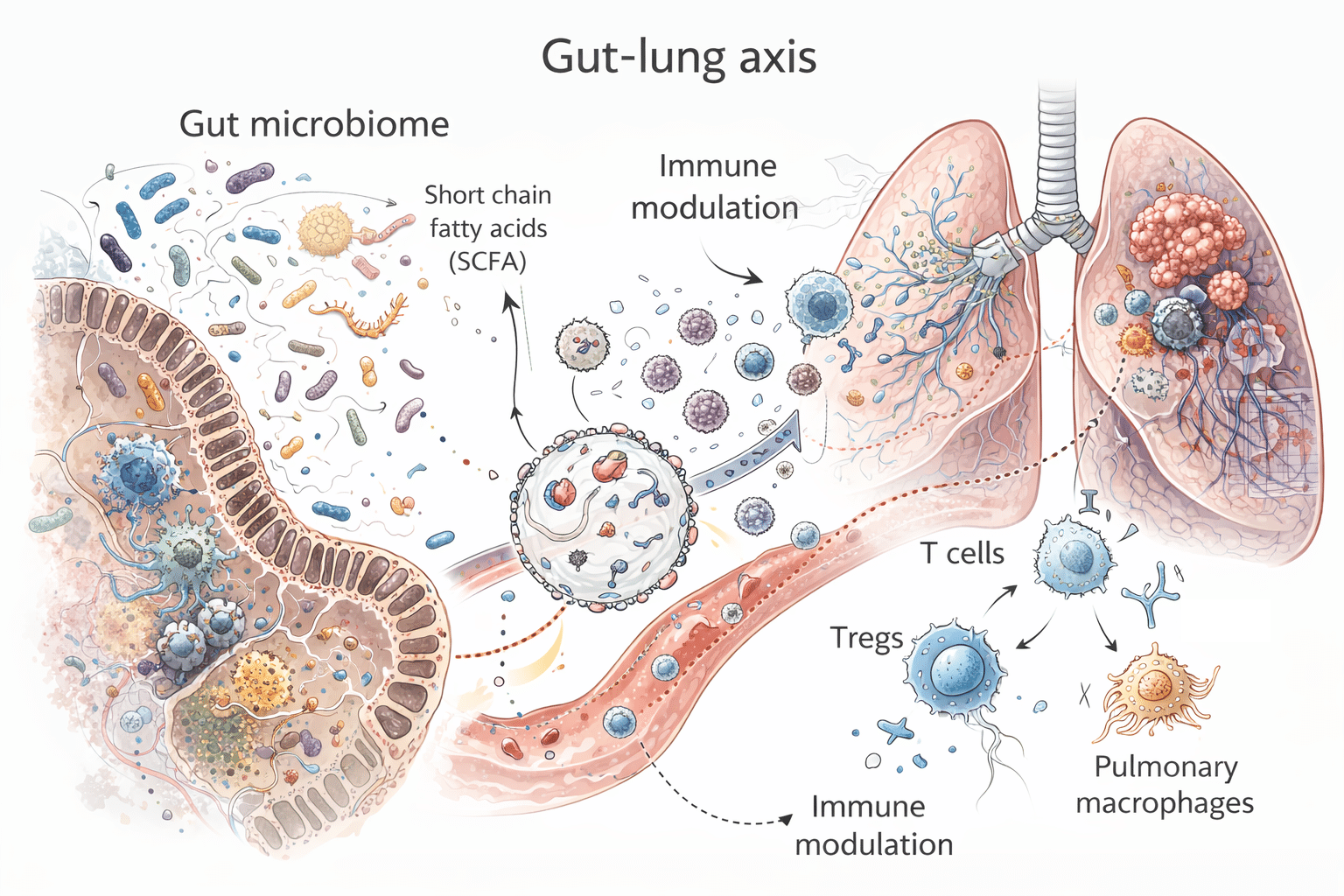In situ mapping of the tumor microenvironment in lung cancer
Lung cancer—most commonly non-small cell lung cancer (NSCLC)—remains one of the leading causes of cancer-related death worldwide. Despite major advances in immunotherapies, especially immune checkpoint inhibitors (ICIs), the effectiveness of current treatments is still limited, and the predictive power of established biomarkers such as PD-L1 expression and tumor mutational burden (TMB) is insufficient. The aim of this research project is to discover new tissue-based biomarkers that can better predict how a patient will respond to immunotherapy.
Over the past decade, the tumor microenvironment (TME) has become a central focus of tumor biology. A growing body of evidence links the intratumoral density of specific immune cell populations to patient survival and treatment response. CD8+ cytotoxic T cells are key effectors of anti-tumor immunity, as they recognize and eliminate tumor cells and can support long-term immune protection. Innate lymphoid cells (ILCs) can regulate the TME both directly and indirectly, while NK cells and γδ T cells can also kill tumor cells directly. In contrast, regulatory T cells (Tregs), myeloid-derived suppressor cells (MDSCs), and M2-like tumor-associated macrophages (TAMs) are generally considered pro-tumor immune populations. TAMs and Tregs can influence CD8+ T-cell infiltration and may promote tumor cell survival and proliferation.
Importantly, T cells can become functionally exhausted in an immunosuppressive TME, meaning that cell density and simple ratios do not always faithfully reflect anti-tumor immune activity. Therefore, immune checkpoints (e.g., PD-L1, CTLA-4) and exhaustion-associated markers expressed on CD8+ T cells (e.g., PD-1, LAG-3, TIM-3, TIGIT) can provide additional, valuable information about the functional state of the tumor immune microenvironment.
The project’s core objectives:
- Construct tissue microarrays (TMAs) from formalin-fixed, paraffin-embedded (FFPE) samples of available retrospective and prospective lung tumor cohorts, including non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC).
- Perform detailed tumor microenvironment (TME) characterization using immunohistochemistry (IHC), RNAscope assays, and multiplex spatial technologies (Akoya PhenoCycler, NanoString GeoMx Digital Spatial Profiler).
- Develop refined scoring frameworks and computational image-analysis workflows, and implement integrated processing of parallel spatial RNA–protein expression data (spatial biology).
- Apply advanced multivariable statistical models and machine learning algorithms to identify prognostic and predictive biomarkers.
The impact of the gut microbiome on anti-tumor immunity and immunotherapy
Lung cancer—especially non-small cell lung cancer (NSCLC)—remains a leading cause of cancer mortality worldwide. Although immune checkpoint inhibitors (ICIs) have transformed first-line treatment for many patients with advanced NSCLC, durable benefit is still achieved only in a subset: 5-year overall survival is around ~30% in biomarker-selected high PD-L1 populations, and typically ~10–20% in less selected groups.
One of the most impactful developments in recent oncology has been the realization that the gut microbiome can modulate anti-tumor immunity, shape the TME, and influence the efficacy of immunotherapy. Beyond gastrointestinal cancers, accumulating evidence also supports clinically meaningful microbiome–immunity links in NSCLC, including mechanisms related to microbial metabolites, immune priming, myeloid programming, and barrier-linked inflammation.
Publications in the topic: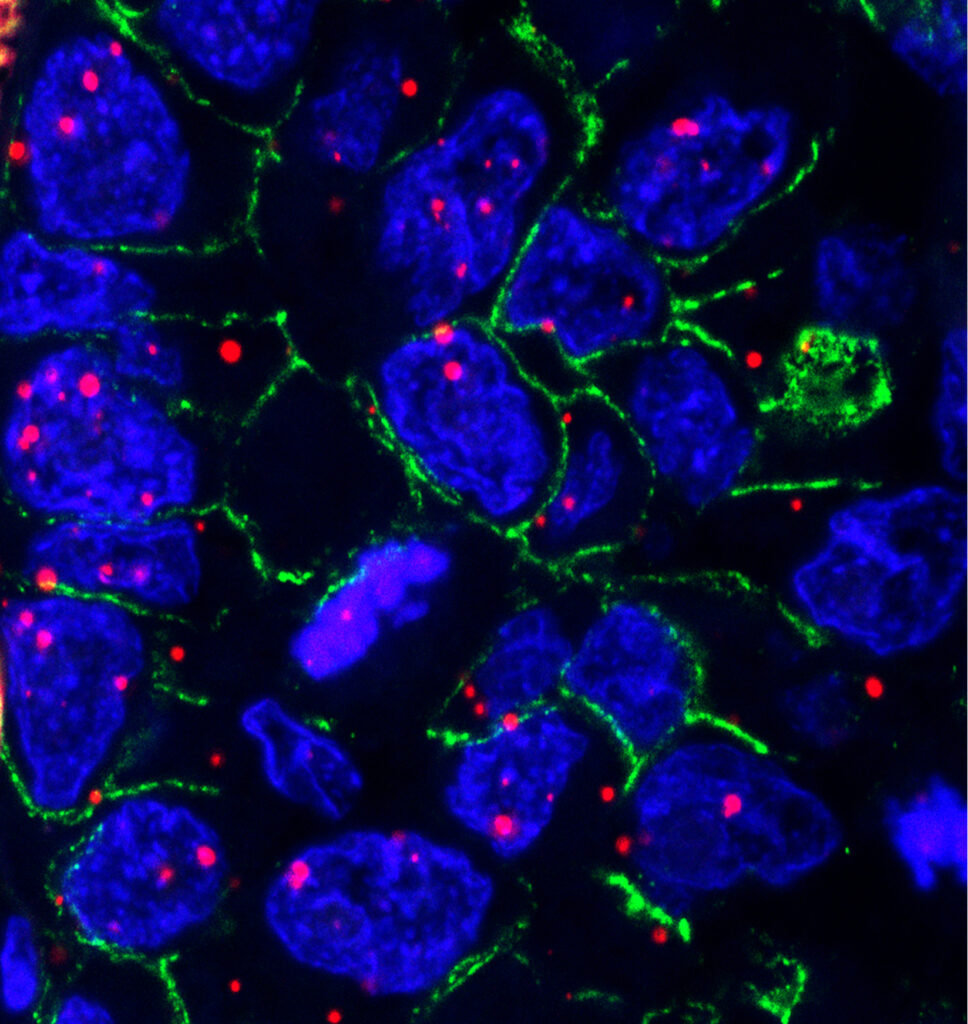
Somodi C, Dora D, Horváth M, Szegvari G, Lohinai Z. Gut microbiome changes and cancer immunotherapy outcomes associated with dietary interventions: a systematic review of preclinical and clinical evidence. JOURNAL OF TRANSLATIONAL MEDICINE. 2025 Jul 8;23(1):756.
Dora D, Revisnyei P, Pasic A, Galffy G, Dulka E, Mihucz A, Roskó B, Szincsak S, Iliuk A, Weiss GJ, Lohinai Z. Host and bacterial urine proteomics might predict treatment outcomes for immunotherapy in advanced non-small cell lung cancer patients. FRONTIERS IN IMMUNOLOGY. 2025 Apr 14;16:1543817.
Dora, David., Kiraly, P., Somodi, C. Balazs, Ligeti; Edit, Dulka; Gabriella, Galffy; Zoltan, Lohinai ✉. Gut metatranscriptomics based de novo assembly reveals microbial signatures predicting immunotherapy outcomes in non-small cell lung cancer. JOURNAL OF TRANSLATIONAL MEDICINE 22, 1044 (2024).
Dora, David ✉ ; Szőcs, Emőke; Soós, Ádám; Halasy, Viktória; Somodi, Csenge; Mihucz, Anna; Rostás, Melinda; Mógor, Fruzsina; Lohinai, Zoltan; Nagy, Nándor From bench to bedside: an interdisciplinary journey through the gut-lung axis with insights into lung cancer and immunotherapy FRONTIERS IN IMMUNOLOGY 15 Paper: 1434804 , 29 p. (2024)
David, Dora ; Balazs, Ligeti ; Tamas, Kovacs ; Peter, Revisnyei ; Gabriella, Galffy ; Edit, Dulka ; Dániel, Krizsán ; Regina, Kalcsevszki ; Zsolt, Megyesfalvi ; Balazs, Dome ✉ ; Zoltan, Lohinai ✉. Non-Small Cell Lung Cancer Patients Treated with Anti-PD1 Immunotherapy Show Distinct Microbial Signatures and Metabolic Pathways According to Progression-free Survival and PD-L1 status ONCOIMMUNOLOGY 12 : 1 Paper: 2204746 , 15 p. (2023)
Dora, David; Weiss, Glen J; Megyesfalvi, Zsolt; Gállfy, Gabriella; Dulka, Edit; Kerpel-Fronius, Anna; Berta, Judit; Moldvay, Judit; Dome, Balazs ✉ ; Lohinai, Zoltan ✉ Computed Tomography-Based Quantitative Texture Analysis and Gut Microbial Community Signatures Predict Survival in Non-Small Cell Lung Cancer. CANCERS 15 : 20 Paper: 5091 , 19 p. (2023)
Dóra, Dávid ✉; Bokhari, Syeda Mahak Zahra; Aloss, Kenan; Takács, Péter; Desnoix, Juliane Zsuzsanna; Szklenárik, György; Hurley, Patrick Deniz; Lohinai, Zoltán ✉ Implication of the Gut Microbiome and Microbial-Derived Metabolites in Immune-Related Adverse Events: Emergence of Novel Biomarkers for Cancer Immunotherapy INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES 24 : 3 Paper: 2769 , 26 p. (2023)
Book chapter:
Dóra, D. A bélidegrendszer: bél-agy tengely In: Tulassay, Zsolt (szerk.) Gasztroenterológia Budapest, Magyarország : Medicina Könyvkiadó (2023) 1,180 p. pp. 14-23. , 10 p.
Key questions we address:
- Which spatially defined TME archetypes (cell density and organization) associate with ICI response and survival?
- How do gut microbiome composition and function correlate with measurable TME features?
- What is the functional state of tumor-infiltrating immune cells (activation/exhaustion), and how does it relate to microbiome patterns?
- Can specific lymphocyte dysfunction phenotypes be linked to taxonomic or metabolic microbiome signatures (gut and/or intratumoral signals)?
- Can microbial biomarkers predict patient benefit from immunotherapy?
Our goal is to identify new prognostic and predictive biomarkers across the TME and the microbiome, and to clarify how gut and intratumoral microbes may shape anti-tumor immunity—creating a rationale for future microbiome-informed adjuvant strategies.
Main goals of the project:
- Link spatial biology data from multiplex tumor microenvironment profiling with the gut microbiome.
- Analyze the intratumoral microbiome and relate it to the gut microbiome, tumor microenvironment features, and patients’ treatment response.
- Use in vitro experiments and mouse studies to validate the statistically identified associations.
- Develop microbial adjuvants to improve immunotherapy, and test them preclinically in tumor mouse models.
Tools and methods
- Immunohistochemistry (IHC), immunocytochemistry (ICC), and immunofluorescence (IF) on frozen and formalin-fixed, paraffin-embedded (FFPE) sections
- RNAscope and RNAscope–IF multiplexing on frozen and FFPE sections
- Standard histology workflow; tissue microarray (TMA) design and participation in TMA construction
- Development and use of advanced microscopic scoring systems; computational image analysis and morphometrics on chromogenic and fluorescently stained sections; confocal microscopy and image analysis. Multimodal spatial biology data analysis using QuPath and ImageJ software suites.
- Microbiome sequencing data analysis (metagenome, metatranscriptome): generating raw abundance matrices, normalization, taxonomic profiling and diversity analyses; metabolic and transcriptomic pathway analysis using MetaPhlAn, HUMAnN, Kraken, and de novo assembly pipelines; data visualization; advanced statistical modeling and machine learning
- Clinical data handling: database building, data management, and quality control
- Mouse experiments: preparation and execution; tissue and organ collection and processing for histology and downstream molecular analyses